Microsatellite Marker Development and Characterization in the Spotted Babylon, Babylonia areolata (Link, 1807): Detection of Duplicated Loci at High Frequency 
* The authors who contribute equally
 Author
Author  Correspondence author
Correspondence author
International Journal of Aquaculture, 2012, Vol. 2, No. 2 doi: 10.5376/ija.2012.02.0002
Received: 15 Mar., 2012 Accepted: 06 Apr., 2012 Published: 14 May, 2012
Zhang et al., 2012, Microsatellite Marker Development and Characterization in the Spotted Babylon, Babylonia areolata (Link, 1807): Detection of Duplicated Loci at High Frequency, International Journal of Aquaculture, Vol.2, No.2 5-10 (doi: 10.5376/ija.2012. 02.0002)
Four polymorphic microsatellite markers were developed for the spotted babylon, Babylonia areolata, from a microsatellite enriched library. Those markers, characterized in 32 individuals from one wild population, were polymorphic with allele numbers ranging from 5 to 15 per locus, expected and observed heterozygosity ranging from 0.60 to 0.92 and from 0.36 to 0.88, respectively. One locus showed significant (p<0.05 after Bonferroni correction) deviation from Hardy–Weinberg equilibrium, probably due to the presence of null alleles. These microsatellite markers should be useful for population genetics studies in this species. In addition, nine primer pairs amplified duplicated microsatellites, with four multiple loci clearly linked in tandem, which provides a good opportunity to study the evolution of repetitive DNA sequences in the genome.
The spotted babylon Babylonia areolata (Link, 1807), also called maculated ivory whelk, is a marine gastropod mollusc distributed in southeastern Asia. Its geographic distribution stretches from Ceylon and Nicobar Islands through the Gulf of Thailand, along Vietnamese and Chinese coasts to Taiwan (Altena and Gittenberger, 1981). It is mostly found on mud and sandy littoral bottoms not exceeding 5~10 m in depth (Panichasuk, 1996). The spotted babylon is a popular model mollusk used in studies on heavy-metal toxicity and biologic poisoning toxins transmission studies (Chen and Chou, 1998; Supanopas et al., 2005; Tanhan et al., 2005) since its being the popular and nutritious seafood. Recent years the wild populations have declined rapidly duo to fast-growing demand for consumption and environmental pollution. The spotted babylon has been a commercially important aquaculture species in Thailand (Chaitanawisuti et al., 2002; Kritsanapuntu et al., 2006) and China (Liang et al., 2005). The individuals from Thailand have greater growth vigor than native ones in Hainan Island, so most hatchery populations in Hainan come from Thailand lines. But little is known about the genetic structure and variation in this species (Hualkasin et al., 2008, Wang et al., 2011a). To support population structure analyses and facilitate stock management (Wang et al., 2010b), we developed a microsatellite enriched library and characterized 4 novel polymorphic microsatellite markers for the spotted babylon. In addition, we found high percentage of duplicated microsatellites clearly linked in tandem, which provides a good opportunity to study the evolution of repetitive DNA sequences in the genome though these multi-locus microsatellites may not be suitable for using as genetic markers because it is often impossible to discern the locus-specific alleles (Zhang, 2004).
1 Results
1.1 Microsatellite sequences
One hundred and twenty-eight positive clones were sequenced, producing a total of 59 252 bp of DNA sequences. Analysis with MISA identified 118 microsatellite-containing sequences, corresponding to an enrichment efficiency of 97.5%. The 118 microsatellite-containing sequences harbored 406 microsatellites with di- (85.5%), tri- (4.2%), tetra- (6.2%), penta- (0.7%), hexa- (1.7%), and hepata- (1.7%) repeats. Ninety (76.3%) sequences contained more than one microsatellite. But most of these sequence contained clustered and complex repeated structures, only fifty-six sequences had sufficient flanking regions for primer designing.
1.2 Amplification and polymorphism
Thirteen primer pairs showed reliable and reproducible amplification in 32 wild individuals. Sequences of these 13 microsatellites were submitted to GenBank (accession numbers FJ594998 – FJ595016). Four of the 13 primer pairs produced no more than 2 alleles per individual and were considered valid single-locus microsatellite markers (Table 1). The 4 single-locus microsatellite loci were polymorphic with allele numbers ranging from 5 to 15 per locus (mean, 11±4.55).
The expected and observed heterozygosity ranged from 0.60 to 0.92 (mean, 0.79±0.14) and from 0.52 to 0.90 (mean, 0.67±0.24), respectively (Table 1). No linkage disequilibrium was detected among all the loci at p>0.05 (after Bonferroni correction). One locus (HUBA07) showed significant (p<0.05 after Bonferroni correction) deviation from Hardy–Weinberg equilibrium. HUBA10 showed signs of null alleles as indicated by Micro-checker (1 000 randomizations) (Table 1), suggesting that the presence of null alleles is the main cause for the deviation from HWE at these loci. The four microsatellite markers developed here should be useful for population genetics studies in this species.
 Table 1 Primer sequences and characteristics of 4 Babylonia areolata single-locus microsatellite markers. The number of individual analyzed (n), number of alleles (a), allele size range in base pairs (as), observed (HO) and expected (HE) heterozygosities are presented for each locus. Ta is the annealing temperature and PHWE is the probability of Hardy–Weinberg equilibrium |
1.3 Duplicated loci
The remaining nine (69%, 9 of 13) duplicated loci amplified more than 2 alleles in most individual (Table 2).
 Table 2 Information of nine Babylonia areolata duplicated loci. The number of individuals analyzed (n), allele size across multi-locus range in base pairs (as) are presented |
The extra alleles can not be eliminated through increasing amplification stringency (lower Mg2+ concentration and higher annealing temperature), suggesting that they were not caused by nonspecific amplification, but by amplification of duplicated loci. Four (30.8%, 4 of 13 ) of them amplified 2 to 4 parallel loci per individual between 100 and 550 bp, which were clearly linked in tandem as they were separated in well-defined patterns. For example, HUBA02 amplified unambiguous non-overlapping two parallel loci linked obviously, one to two fragments that differed by 8n bp per locus, with a locus-to-locus distance of about 145 bp (Figure 1). This can be explained by the model of [forward primer] - 145 bp- [forward primer] -267 bp - (8 bp) n- [reverse primer]. This finding implies that one of the primers sequence may be located within a satellite near the microsatellite loci so that there is significant tandem duplication of microsatellite loci and/or their flanking sequences in B. areolata genome.
2 Discussion
Those 4 duplicated loci amplified parallel loci separated in well-defined pattern observed in this study were similar to those microsatellite loci that were linked in tandem with duplicated structures, e.g. satellites or mini-satellite sequences found in Atlantic surfclam, Spisula solidissima (6.7%, 2 of 30) (Wang et al., 2009b), white-lipped pearl oyster, Pinctada maxima (11.1%, 2 of 18) (Wang et al., 2009a), and hard clam Mercenaria mercenaria (21.2%, 11/52) (Wang et al., 2010a). As far as we can determine, high frequency of duplicated locus linked in tandem such as the 30.8% observed in this study has not been reported in any mollusk. This finding implies that the complex duplications of B. areolata genome are partly characterized by overlapping and/or linkage in tandem of microsatellites, mini-satellites and satellites.
There probably is high frequency of microsatellite sequences in B. areolata genomes, since we obtained an enrichment efficiency of 97.5% (118 of 121) in this study, which is much higher than that obtained in other four molluscan species whose microsatellite markers were developed in parallel in our laboratory using the same protocol and reagents: 63.3% for Spisula solidissima (Wang et al., 2009b), 53.2% for Pinctada maxima (Wang et al., 2009a), 59.4% for Mercenaria mercenaria (Wang et al., 2010a), and 58.5% for Haliotis diversicolor (Wang et al., 2011c). Extraordinarily, there are sixty-two microsatellite sequences (52.5%, 62 out of 118) contained too long and complex microsatellites to have sufficient flanking regions for primer designing, such high frequency was also reported in B. areolata (Chen et al., 2009; Wang et al., 2011a) and other whelk e.g. Buccinum undatum (Weetman et al., 2005). These results suggested that the B. areolata genome may be a good model to study the evolution of genome duplication in mollusk.
The 4 polymorphic microsatellite markers developed here are useful addition to the collection of other molecular markers that are now available for B. areolata, including mitochondrial markers and other microsatellites markers (Chen et al., 2009; Wang et al., 2011a). They will prove valuable for future population genetic studies and in tracking of hatchery strains in this species.
The finding of significant proportions of locus duplication suggests that the use and isolation of microsatellite markers in B. areolata may be challenging. Perhaps further efforts in this species should be directed to the development of SNP markers based on the second generation sequencing technology.
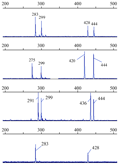 Figure 1 PCR amplifying fragments of duplicated loci HUBA02 in 4 Babylonia areolata individuals shown by GeneMapper 3.5 software |
3 Materials and Methods
3.1 Samples and DNA extraction
Thirty-two B. areolata individuals were collected from a wild population at the seashore near Qiaogang town of Beihai city, Guanxi Province, China. DNA was extracted from the ethanol preserved gastropod muscles using the Cell/Tissue Genomic DNA extraction kit (TianGen, Beijing, China) (Lu et al., 2011).
3.2 Microsatellite-enriched library construction and sequencing
A microsatellite-enriched DNA library was constructed using a selective hybridization and magnetic bead enrichment protocol as described in Wang et al (2010a). Briefly, about 38 µg genomic DNA of three individuals was digested by the restriction enzyme Mbo I (Takara, Dalian, China). Fragments of 300~1 000 bp collected with the Gel DNA Extraction Kit (TianGen) were ligated to double stranded Mbo I adapters by incubating with T4 DNA ligase (Takara) at 16℃ overnight. Excess adapters were removed by washing with 0.1×TE buffer (pH 8.0) on an Ultrafree column (Pall, CA, USA). DNA fragments with adapters were amplified with linker B for 5 PCR cycles and purified using an Ultrafree column (Pall). The amplified products were denatured and then hybridized to biotin-labeled (CA)12, (GA)12, (ACA)8, (AGA)8, (GACA)6 and (GATA)6 oligonucleotides (mixed in advance at the ratio of 3:1:1:1:2:2, total 150 pmol) in 0.5×SSC at 68℃ for 60 min. Washed four times in 0.1×SSC at room temperature, DNA fragments bound to these probes were then captured with Streptavidin MagneSphere® Paramagnetic Particles (Promega, USA) and eluted by DNase-free water. The microsatellite-enriched elution was amplified and purified as described above. The amplified products were cloned using the T-Easy system (Promega). Clones containing potential microsatellite loci were selected and sequenced on ABI 3730xl DNA analyzers at Sangon Biological Engineering Technology & Services Co., Ltd. (Shanghai, China).
3.3 Primer design and genotyping
DNA sequences removed the linkers and T vector were searched against each other and against 17 spotted babylon microsatellite sequences from GenBank (accessed March 19, 2012) using Vector NTI Advance 11.0.0 (http://www.invitrogen.com) to check for duplicates. Microsatellite sequences containing at least 6 di-, 5 tri-, 5 tetra-, 4 penta-, and 3 hexa-, hepta- and octanucleotide repeats were selected using MISA software (http://pgrc.ipk-gatersleben.de/misa/). Good sequences with sufficient flanking regions were used for primer design with Primer3 (http://biotools.umassmed.edu/bioapps/primer3_www.cgi). An M13 (-21) universal leading sequence (5-TGTAAAACGACGGCCAGT-3) was added to the 5' end of each forward primer (Schuelke, 2000), and primers were synthesized by Sangon Biological Engineering Technology & Services Co., Ltd (Shanghai, China).
All primer pairs were tested for PCR amplification in the 32 wild individuals. PCR was conducted in a 10 mL solution containing about 30 ng template DNA, and reagents (TianGen) as follows: 1× reaction buffer containing 20 mM Tris-Hcl (pH 8.4), 20 mM KCl and 10 mM (NH4)2SO4, 1.2~2.0 mM MgCl2, 0.2 mM each dNTP, 0.4 units Taq polymerase, 0.2 pmol M13 (-21) tailed forward primer, 0.6 pmol reverse primer, 0.6 pmol M13 (-21) primer labeled with fluorescent dyes (FAM, VIC, NED or PET, Applied Biosystems, Foster City, CA). PCR was conducted on a Master Cycler Gradient (Eppendorf, Germany) with the following steps: 94℃ for 4 min followed by 34 cycles of 94℃ for 30 sec, 30 sec annealing (temperatures indicated in Table 1) for 45 s, followed by 10 cycles of 94℃ for 30 sec, 53℃ for 45 sec, 72℃ for 45 sec, and a final extension at 72℃ for 10 min. Amplified products were detected and sized on an ABI 3130 Prism Genetic Analyzer (Applied Biosystems) using GS-500LIZ (Applied Biosystems) as the size standard. Allele scoring was performed with GeneMapper v3.5 (Applied Biosystems). Samples failing to amplify at the first time were re-amplified once.
3.4 Statistical analysis
The Micro-Checker 2.2.1 software (van Oosterhout et al., 2004) was used to identify possible genotyping errors and null alleles (1 000 randomizations). GENEPOP on the Web (http://genepop.curtin.edu.au/) was used to test deviations from Hardy-Weinberg equilibrium (HWE) for each locus and linkage disequilibrium between all pairs (exact tests, 1 000 iterations). ARLEQUIN 3.0 software (Excoffier et al., 2005) was used to calculate the observed (HO) and expected (HE) heterozygosity. All tests were corrected for multiple comparisons by Bonferroni’s correction (Rice, 1989).
Author Contributions
NZ and YQ are the executor of experimental research in this study, they have the same contribution to the paper. YW and XZH make the experimental design, data analysis, paper writing and revising. AMW is the supervisor of the project and XFC provides the necessary facilities for the research to be performed.
Acknowledgement
This study was supported by the National Natural Science Foundation of China (40866003 and 31060354), the Natural Science Foundation of Hainan (80616, 311024), and the National Basic Research Program of China (973 program, 2010CB126405).
Reference
Altena C.O.V.R., and Gittenberger E., 1981, The genus Babylonia (Prosobranchia, Buccinidae), Zoologische Verhandelingen, 188: 1-57
Chaitanawisuti N., Kritsanapuntu S., and Natsukari Y., 2002, Economic analysis of a pilot commercial production for spotted babylon Babylonia areolata Link, 1807 marketable sizes using a flow-through culture system in Thailand, Aquaculture Res., 33: 1-8
http://dx.doi.org/10.1046 /j.1365 2109.2002.00790.x
Chen C.Y., and Chou H.N., 1998, Transmission of the paralytic shellfish poisoning toxins, from dinoflagellate to gastropod, Toxicon, 36: 515-522
http://dx.doi.org/10.1016/S0041-0101(97)00093-7
Chen F., Ke C.H., Wang D.X., Chen J., Zhan X., and Luo X., 2009, Isolation and characterization of microsatellite loci in Babylonia areolata and cross-species amplification in Babylonia formosae habei, Permanent Genetic Resources, 9: 661-663
Excoffier L., Laval G., and Schneider S., 2005, Arlequin ver. 3.0: an integrated software package for population genetics data analysis, Evol Bioinform Online, 1: 47-50
Hualkasin W., Tongchuai W., Chotigeat W., and Phongdara A., 2008, Phylogeography of Ivory shell (Babylonia areolata) in the Gulf of Thailand revealed by COI gene structure and differentiation of shell color by ITS1 DNA, Songklanakarin J. Sci. Technol, 30 (2): 141-146
Kritsanapuntu S., Chaitanawisuti N., Santhaweesuk W., and Natsukari Y., 2006, Growth, production and economic evaluation of earthen ponds for monoculture and polyculture of juveniles spotted babylon (Babylonia areolata) to marketable sizes using large-scale operation, Journal of Shellfish Research, 25(3): 913-918
Liang F., Mao Y., Yu X., and Liu H., 2005, Experiment on artificial breeding of Babylonia areolata, Transactions of Oceanology and Limnology, 2005(1): 79-85
Hu Lu, Xuzhen Huang, Jintian Zhu, Yan Wang, 2011, Chromosomal Mapping of Ribosomal rRNA Genes in the Small Rock Oyster, Saccostrea mordax (Gould 1850), International Journal of Marine Science, 1(2): 2-5
http://dx.doi.org/10.5376/ijms.2011.01.0002
Panichasuk P., 1996, Areola babylon, Babylonia areolata Link, 1807, Thai Fish Gazette, 49: 107-117
Rice W.R., 1989, Analyzing tables of statistical tests, Evolution, 43: 223-225
http://dx.doi.org/10.2307/2409177
Schuelke M., 2000, An economic method for the fluorescent labeling of PCR fragments, Nat Biotechnol, 18: 233-234
http://dx.doi.org/10.1038/72708 PMid:10657137
Supanopas P., Sretarugsa P., Kruatrachue M., Pokethitiyook P., and Upatham E.S., 2005, Acute and subchronic toxicity of lead to the spotted babylon, Babylonia areolata (neogastropoda, buccinidae), Journal of Shellfish Research, 24: 91-98
Tanhan P., Sretarugsa P., Pokethitiyook P., Kruatrachue M., and Upatham E.S., 2005, Histopathological alterations in the edible snail, Babylonia areolata (spotted babylon), in acute and subchronic cadmium poisoning, Environ Toxicol, 20: 142-149
http://dx.doi.org/10.1002/tox.20088 PMid:15793826
van Oosterhout C., William F.H., Derek P.M.W., and Peter S., 2004, MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data, Molecular Ecology Notes, 4: 535-538
http://dx.doi.org/10.1111/j.1471-8286.2004.00684.x
Wang Y., Liu N., Shi Y., Gu Z., and Wang A., 2009a, Development and characterization of sixteen new microsatellite markers for the white-lipped pearl oyster, Pinctada maxima (Jameson, 1901), Molecular Ecology Resource, 9: 1460-1466
Wang Y., Lu H., Zheng J., Long K., Shi Y.H., Gu Z.F., and Wang A.M., 2011a, Eight polymorphic microsatellite markers for the spotted babylon, Babylonia areolata (Buccinidae), Genet Mol Res, 10: 3230-3235
http://dx.doi.org/10.4238/2011.December.21.5 PMid:22194180
Wang Y., Wang A., and Guo X., 2009b, Development and characterization of 30 polymorphic microsatellite markers for the Atlantic surfclam, Spisula solidissima (Dillwyn, 1817), Molecular Ecology Resource, 9: 1264-1267
http://dx.doi.org/10.1111/j.1755-0998.2009.02660.x PMid:21564897
Wang Y., Wang A., and Guo X., 2010a, Development and Characterization of Polymorphic Microsatellite Markers for the Northern Quahog Mercenaria mercenaria (Linnaeus, 1758), Journal of Shellfish Research, 29: 77-82
http://dx.doi.org/10.2983/035.029.0130
Wang Y., Wang F., Shi Y.H., Gu Z.F., and Wang A.M., 2011b, Development and characterization of 60 microsatellite markers in the abalone Haliotis diversicolor, Genet Mol Res, 10: 860-866
http://dx.doi.org/10.4238/vol10-2gmr1182 PMid:21574142
Wang Y., Wang X., Wang A., and Guo X., 2010b, A 16-microsatellite multiplex assay for parentage assignment in the eastern oyster (Crassostrea virginica Gmelin), Aquaculture, 308: S28-S33
http://dx.doi.org/10.1016/j.aquaculture.2010.05.037
Weetman D., Hauser L., Shaw P.W., and Bayes M., 2005, Microsatellite markers for the whelk Buccinum undatum, Mol Ecol Notes, 5: 361-362
http://dx.doi.org/10.1111/j.1471-8286.2005.00926.x
Zhang D.X., 2004, Lepidopteran microsatellite DNA: redundant but promising, Trends in Ecology & Evolution, 19: 507-509
http://dx.doi.org/10.1016/j.tree.2004.07.020
. PDF(461KB)
. FPDF(win)
. HTML
. Online fPDF
Associated material
. Readers' comments
Other articles by authors
. Nan Zhang
. Ying Qiu
. Xuzhen Huang
. Xuefen Chen
. Aimin Wang
. Yan Wang
Related articles
. Babylonia areolata
. Microsatellites
. Duplicated loci
. Population genetics
. Fishery
. Gastropod
. Spotted babylon
Tools
. Email to a friend
. Post a comment


